Seed labeling with scANVI#
In this tutorial, we go through the steps of training scANVI for seed annotation. This is useful for when we have ground truth labels for a few cells and want to annotate unlabelled cells. For more information, please refer to the original scANVI publication.
Plan for this tutorial:
Loading the data
Creating the seed labels: groundtruth for a small fraction of cells
Training the scANVI model: transferring annotation to the whole dataset
Visualizing the latent space and predicted labels
Note
Running the following cell will install tutorial dependencies on Google Colab only. It will have no effect on environments other than Google Colab.
!pip install --quiet scvi-colab
from scvi_colab import install
install()
WARNING: Running pip as the 'root' user can result in broken permissions and conflicting behaviour with the system package manager. It is recommended to use a virtual environment instead: https://pip.pypa.io/warnings/venv
import tempfile
import numpy as np
import scanpy as sc
import scvi
import seaborn as sns
import torch
scvi.settings.seed = 0
print("Last run with scvi-tools version:", scvi.__version__)
Last run with scvi-tools version: 1.1.0
Note
You can modify save_dir below to change where the data files for this tutorial are saved.
sc.set_figure_params(figsize=(6, 6), frameon=False)
sns.set_theme()
torch.set_float32_matmul_precision("high")
save_dir = tempfile.TemporaryDirectory()
%config InlineBackend.print_figure_kwargs={"facecolor": "w"}
%config InlineBackend.figure_format="retina"
Data Loading#
For the purposes of this notebook, we will be labeling 4 cell types in a dataset of purified peripheral blood mononuclear cells from 10x Genomics:
CD4 Regulatory T cells
CD4 Naive T cells
CD4 Memory T cells
CD8 Naive T cells
adata = scvi.data.purified_pbmc_dataset(
save_path=save_dir.name,
subset_datasets=["regulatory_t", "naive_t", "memory_t", "naive_cytotoxic"],
)
INFO Downloading file at /tmp/tmp43gcrr_r/PurifiedPBMCDataset.h5ad
Downloading...: 157054it [00:01, 100788.45it/s]
From now on, we assume that cell type information for each cell is unavailable to us, and we seek to retrieve it.
Automatic annotation using seed labels#
In this section we hand curate and select cells which will serve as our ground truth labels.
We start by putting together a list of curated marker genes, from which we aim at identifying our ground truth cell types. These are extracted from the scANVI publication.
gene_subset = [
"CD4",
"FOXP3",
"TNFRSF18",
"IL2RA",
"CTLA4",
"CD44",
"TCF7",
"CD8B",
"CCR7",
"CD69",
"PTPRC",
"S100A4",
]
Then, we assign a score to every cell as a function of its cell type signature. In order to compute these scores, we need to normalize the data. Because this is not the case of scVI or scANVI, we proceed with a copy of the dataset for this step.
normalized = adata.copy()
sc.pp.normalize_total(normalized, target_sum=1e4)
sc.pp.log1p(normalized)
normalized = normalized[:, gene_subset].copy()
sc.pp.scale(normalized)
Now, here we have two helper functions that will help in scoring the cells, as well as taking the most confident cells with respect to these scores.
def get_score(normalized_adata, gene_set):
"""Returns the score per cell given a dictionary of + and - genes
Parameters
----------
normalized_adata
anndata dataset that has been log normalized and scaled to mean 0, std 1
gene_set
a dictionary with two keys: 'positive' and 'negative'
each key should contain a list of genes
for each gene in gene_set['positive'], its expression will be added to the score
for each gene in gene_set['negative'], its expression will be subtracted from its score
Returns
-------
array of length of n_cells containing the score per cell
"""
score = np.zeros(normalized_adata.n_obs)
for gene in gene_set["positive"]:
expression = np.array(normalized_adata[:, gene].X)
score += expression.flatten()
for gene in gene_set["negative"]:
expression = np.array(normalized_adata[:, gene].X)
score -= expression.flatten()
return score
def get_cell_mask(normalized_adata, gene_set):
"""Calculates the score per cell for a list of genes, then returns a mask for
the cells with the highest 50 scores.
Parameters
----------
normalized_adata
anndata dataset that has been log normalized and scaled to mean 0, std 1
gene_set
a dictionary with two keys: 'positive' and 'negative'
each key should contain a list of genes
for each gene in gene_set['positive'], its expression will be added to the score
for each gene in gene_set['negative'], its expression will be subtracted from its score
Returns
-------
Mask for the cells with the top 50 scores over the entire dataset
"""
score = get_score(normalized_adata, gene_set)
cell_idx = score.argsort()[-50:]
mask = np.zeros(normalized_adata.n_obs)
mask[cell_idx] = 1
return mask.astype(bool)
We run those function to identify highly confident cells, that we aim at using as seed labels
# hand curated list of genes for identifying ground truth
cd4_reg_geneset = {
"positive": ["TNFRSF18", "CTLA4", "FOXP3", "IL2RA"],
"negative": ["S100A4", "PTPRC", "CD8B"],
}
cd8_naive_geneset = {"positive": ["CD8B", "CCR7"], "negative": ["CD4"]}
cd4_naive_geneset = {
"positive": ["CCR7", "CD4"],
"negative": ["S100A4", "PTPRC", "FOXP3", "IL2RA", "CD69"],
}
cd4_mem_geneset = {
"positive": ["S100A4"],
"negative": ["IL2RA", "FOXP3", "TNFRSF18", "CCR7"],
}
cd4_reg_mask = get_cell_mask(
normalized,
cd4_reg_geneset,
)
cd8_naive_mask = get_cell_mask(
normalized,
cd8_naive_geneset,
)
cd4_naive_mask = get_cell_mask(
normalized,
cd4_naive_geneset,
)
cd4_mem_mask = get_cell_mask(
normalized,
cd4_mem_geneset,
)
seed_labels = np.array(cd4_mem_mask.shape[0] * ["Unknown"])
seed_labels[cd8_naive_mask] = "CD8 Naive T cell"
seed_labels[cd4_naive_mask] = "CD4 Naive T cell"
seed_labels[cd4_mem_mask] = "CD4 Memory T cell"
seed_labels[cd4_reg_mask] = "CD4 Regulatory T cell"
adata.obs["seed_labels"] = seed_labels
We can observe what seed label information we have now
adata.obs.seed_labels.value_counts()
seed_labels
Unknown 42719
CD4 Reg 50
CD4 Nai 50
CD4 Mem 50
CD8 Nai 50
Name: count, dtype: int64
As expected, we use 50 cells for each cell type!
Transfer of annotation with scANVI#
As in the harmonization notebook, we need to register the AnnData object for use in scANVI. Namely, we can ignore the batch parameter because those cells don’t have much batch effect to begin with. However, we will give the seed labels for scANVI to use.
scvi.model.SCVI.setup_anndata(adata, batch_key=None, labels_key="seed_labels")
scvi_model = scvi.model.SCVI(adata, n_latent=30, n_layers=2)
scvi_model.train(100)
Epoch 100/100: 100%|██████████| 100/100 [03:50<00:00, 2.30s/it, v_num=1, train_loss_step=1.75e+3, train_loss_epoch=1.76e+3]
Now we can train scANVI and transfer the labels!
scanvi_model = scvi.model.SCANVI.from_scvi_model(scvi_model, "Unknown")
scanvi_model.train(25)
INFO Training for 25 epochs.
Epoch 25/25: 100%|██████████| 25/25 [02:05<00:00, 5.01s/it, v_num=1, train_loss_step=1.77e+3, train_loss_epoch=1.74e+3]
Now we can predict the missing cell types, and get the latent space
SCANVI_LATENT_KEY = "X_scANVI"
SCANVI_PREDICTIONS_KEY = "C_scANVI"
adata.obsm[SCANVI_LATENT_KEY] = scanvi_model.get_latent_representation(adata)
adata.obs[SCANVI_PREDICTIONS_KEY] = scanvi_model.predict(adata)
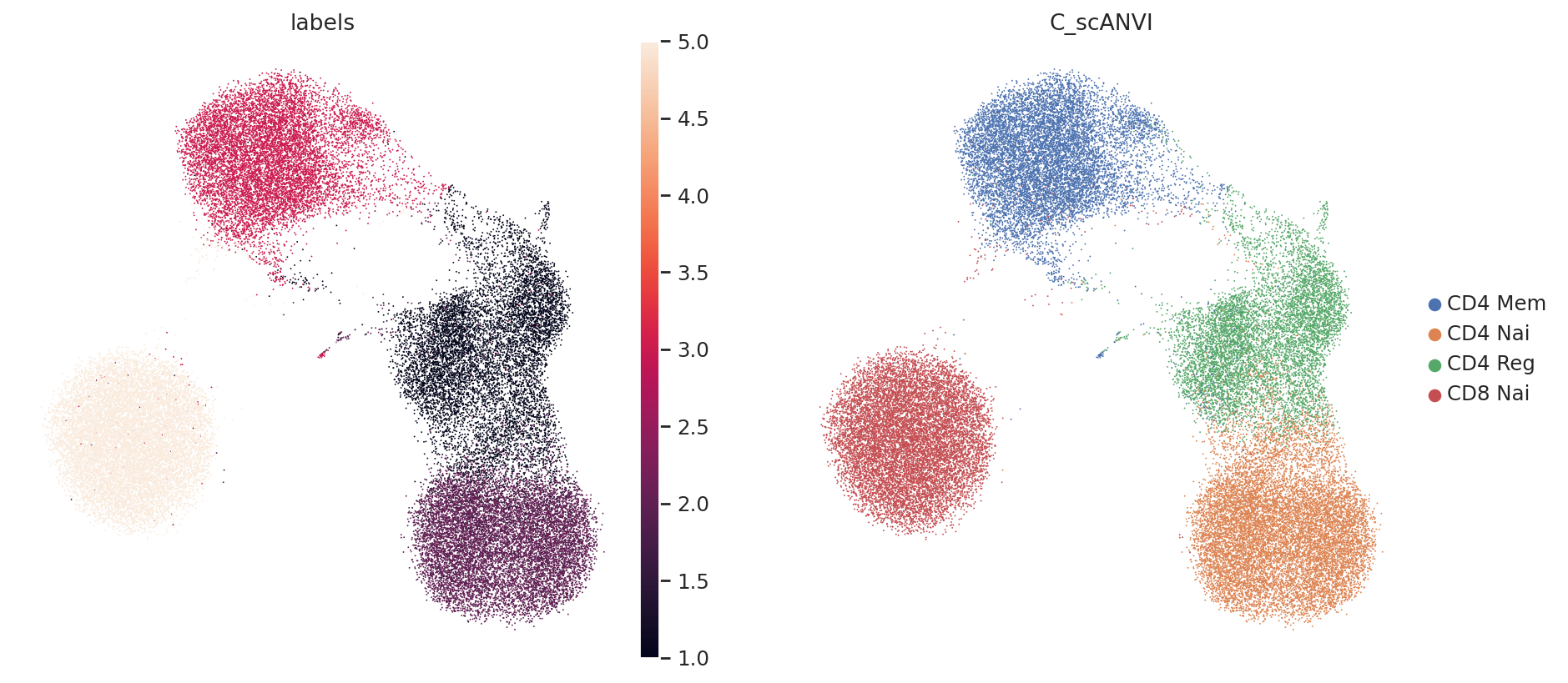
Again, we may visualize the latent space as well as the inferred labels
sc.pp.neighbors(adata, use_rep=SCANVI_LATENT_KEY)
sc.tl.umap(adata)
From this, we can see that it is relatively easy for scANVI to separate the CD4 T cells from the CD8 T cells (in latent space, as well as for the classifier). The regulatory CD4 T cells are sometimes missclassified into CD4 Naive, but this stays a minor phenomenon. Also, we expect that better results may be obtained by careful hyperparameter selection for the classifier. Learn about all of this in our documentation.